KRAS G12D TCR (Clone 9c) CD8+ NFAT-Luciferase Reporter Jurkat Cell Line
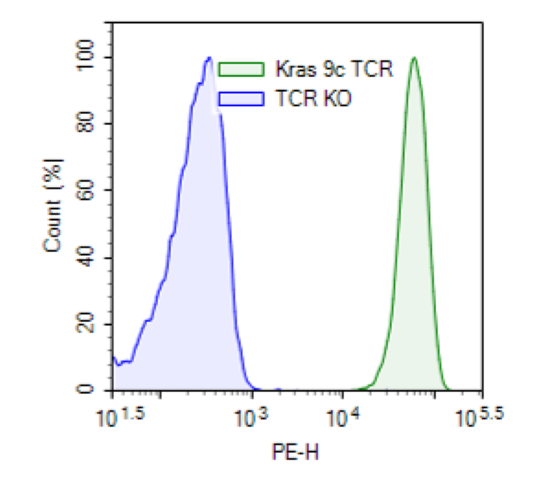
KRAS G12D TCR (Clone 9c) CD8+ NFAT-Luciferase Reporter Jurkat Cell Line was generated from T Cell Receptor (TCR) Knockout NFAT Luciferase Reporter Jurkat Cell Line (BPS Bioscience #78556) by overexpression of human CD8 (NM_001768.6) and KRAS (Kirsten sat sarcoma virus) G12D TCR (Clone 9c) using lentiviral transduction (with CD8a Lentivirus #78648 and KRAS G12D Specific TCR Lentivirus (Clone 9c) #78936). This TCR specifically recognizes the antigen KRAS G12D Peptide, amino acids 10-18 (GADGVGKSA, 9mer).
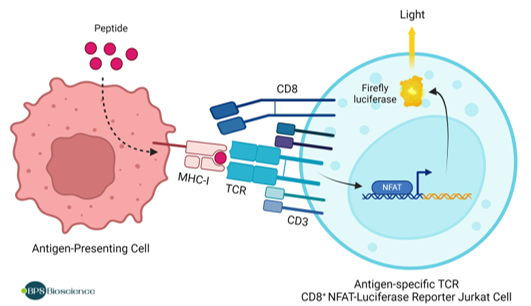
Figure 1: Illustration of the functional co-culture assay used to validate the KRAS G12D TCR (Clone 9c) CD8+ NFAT-Luciferase Reporter Jurkat Cell Line.
Purchase of this cell line is for research purposes only; commercial use requires a separate license. View the full terms and conditions.
| Name | Ordering Information |
| Thaw Medium 2 | BPS Bioscience #60184 |
| Growth Medium 2U | BPS Bioscience #82309 |
| Assay Medium 2D | BPS Bioscience #78755 |
| CD8+ TCR Knockout NFAT-Luciferase Reporter Jurkat Cell Line | BPS Bioscience #78757 |
| HLA-C*08:02 K562 Cell line | BPS Bioscience #78974 |
| KRAS G12D Peptide (10-18, 9mer) | BPS Bioscience #78967 |
| KRAS G12D Peptide (10-19, 10mer) | BPS Bioscience #78969 |
| PE anti-mouse TCR β chain Antibody | BioLegend #109207 |
| ONE-Step™ Luciferase Assay System | BPS Bioscience #60690 |
| 96-well tissue culture plate, white, clear bottom |
The cell line has been screened to confirm the absence of Mycoplasma species.
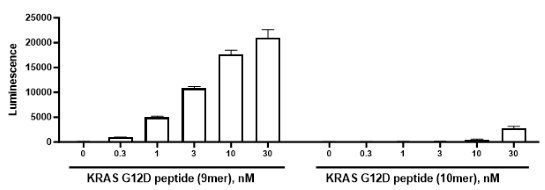
KRAS (Kirsten rat sarcoma virus) are GTPase proteins. They cycle between a GDP-bound inactive state and a GTP-bound (active) form, in a process regulated by two accessory proteins: GEF (guanine exchange factors) and GAPs (GTPase activating proteins). Once activated KRAS can bind to its effectors and regulate multiple signaling pathways, such as the RAF (rapidly accelerated fibrosarcoma)-MEK (mitogen activated protein kinase)-ERK (extracellular regulated kinase) or the PI3K (phosphoinositide 3-kinase)-AKT (protein kinase B)-mTOR (mammalian target of rapamycin) signaling pathways. KRAS mutations account for about 85% of all RAS mutations and are considered one of the main drivers of human cancer, such as in PDAC (pancreatic ductal adenocarcinoma). One of the amino acids frequently mutated is glycine 12, with the most common form being G12D. Since KRAS are intracellular proteins, they are not amenable to CAR (chimeric antigen receptor)-T cell-based therapies, and the development of inhibitors has also proved challenging. One strategy involves the use of TCR (T cell receptor)-T cells, targeting this antigen. Specific TCR clones have been identified, with a KRAS G12D-specific TCR (clone 9c) preferentially being reactive against KRAS G12D peptide (10-18, 9mer), in comparison with KRAS G12D peptide (10-19, 10mer) and being unable to recognize the wild-type KRAS peptides. On the other hand, a KRAS G12D-specific TCR (clone 10) is preferentially reactive against KRAS G12D peptide (10-19, 10mer), in comparison with KRAS G12D peptide (10-18, 9mer) and it also does not recognize wild-type KRAS peptides. Results from a trial using a KRAS G12D HLA-C*08:02 restricted TCR demonstrated the potential of this approach for the treatment of PDAC. The use of neoantigen specific TCR-T cells, targeting single amino acid mutations, is thus an exciting and promising cancer therapy.
CD8 (Cluster of Differentiation 8) is a co-receptor of TCR and a typical marker of cytotoxic T cells. The TCR protein complex is found on the surface of T cells and is reponsible for recognizing antigens bound to MHC (Major Histocompatibility Complex) molecules. Stimulation of the TCR results in the activation of downstream NFAT (Nuclear factor of Activated T-cells) transcription factors that induce the expression of various cytokines such as a interleukin-2 to 4, and TNF-alpha. The use of engineered TCR allows T cells to target specific antigens present in cancer cells via the MHC, expanding the portfolio of antigens that can be targeted in cancer cell therapy.
Eric T., et al., 2016 N Engl J Med 375:2255-2262.
Leidner R., et al., 2022 N Engl J Med 386:2112-2119.