PARP Assays
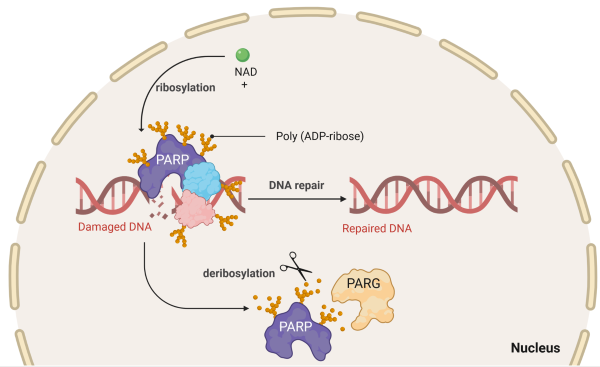
Poly(ADP-ribose) polymerase (PARP) proteins play a crucial role in DNA repair mechanisms within cells. These enzymes are involved in the repair of single-strand DNA breaks through the base excision repair pathway. Inhibition of PARP proteins has emerged as a promising strategy in cancer therapy, particularly in tumors with defective DNA repair mechanisms, such as those with BRCA mutations. When PARP is inhibited, it prevents the repair of single-strand breaks, leading to the accumulation of double-strand breaks during DNA replication. In cells with deficient homologous recombination, often seen in certain cancer types, these double-strand breaks become more lethal. Exploiting this vulnerability has led to the development of PARP inhibitors, which selectively target cancer cells with impaired DNA repair pathways. The continued development of new PARP inhibitors with improved selectively and reduced side effects is critical for effective precision medicine in oncology.
A Unique Suite of Assay Kit and Service Options for Your Discovery
BPS Bioscience has the largest portfolio of commercially available kits and inhibitor screening and profiling services for PARP proteins. Our in-house developed kits, available across seven different formats, provide high quality, clear, and consistent results for determining potent PARP inhibitors. Our off-the-shelf assay kits are available as services where we test your molecules in our assays, providing convenient and timely results. We also provide services for a few additional targets not available as assay kit products. To explore the advantages of our different assay formats, visit our Assay Kits page.
| ELISA Chemiluminescent |
ELISA Colorimetric | AlphaLISA | FP trapping on DNA** | FP competitive binding | Fluorogenic | ELISA cell based | PROTAC Optimization | ||||
| Size (reactions) |
96 | 384 | 96 | 384 | 96 | 384 | 96 | 96 | 384 | 96 | 384 |
| PARP1 | 80551 | 80569 | 80580 | 78438 | 80584-1 | 80584-2 78317 |
82293 | 82123* | 78441 (new) | ||
| PARP2 | 80552-1 | 80552-2 | 80581 | 78572 | 78296-1 | 78296-2 78317 |
82294 | 82123* | |||
| PARP3 | 80553-1 | 80553-2 | 78491 | ||||||||
| PARP4 | Available for Screening | ||||||||||
| PARP5A (TNKS1) | 78405-1 | 78405-2 | 78576 | 78489 | |||||||
| PARP5B (TNKS2) | 78406-1 | 78406-2 | 78577 | 78490 | |||||||
| PARP6 | 80556 | ||||||||||
| PARP7 | 79729-1 78814-1 (new) |
79729-2 78814-2 (new) |
|||||||||
| PARP8 | Available for Screening | ||||||||||
| PARP10 | 80560 | ||||||||||
| PARP11 | 80561 | 78492 | |||||||||
| PARP12 | 78504 | ||||||||||
| PARP14 | 80568 | Available for Screening | |||||||||
| PARP15 | 80567 | ||||||||||
| PARP15FL | 78596 | ||||||||||
| PARP16 | Available for Screening | ||||||||||
| PARG | 78858-1 | 78858-2 | 82123* | ||||||||
| ARH3 | 82158 | ||||||||||
| PARylation | 82123* | ||||||||||
FP = Fluorescence Polarization
*82123 is our LysA™ Universal PARylation Assay Kit designed to measure and quantify the amount of total poly-ADP-ribosylation present in cell extracts. This kit is sufficiently sensitive to measure effects of known PARP1, PARP2, and PARG inhibitors.
**PARPtrap™ is our proprietary assay designed to measure DNA complex formations with PARP1 and PARP2 enzymes using fluorescence polarization.
Browse our PARP Screening & Profiling Services and Request QuoteEnzymatic Activity by ELISA
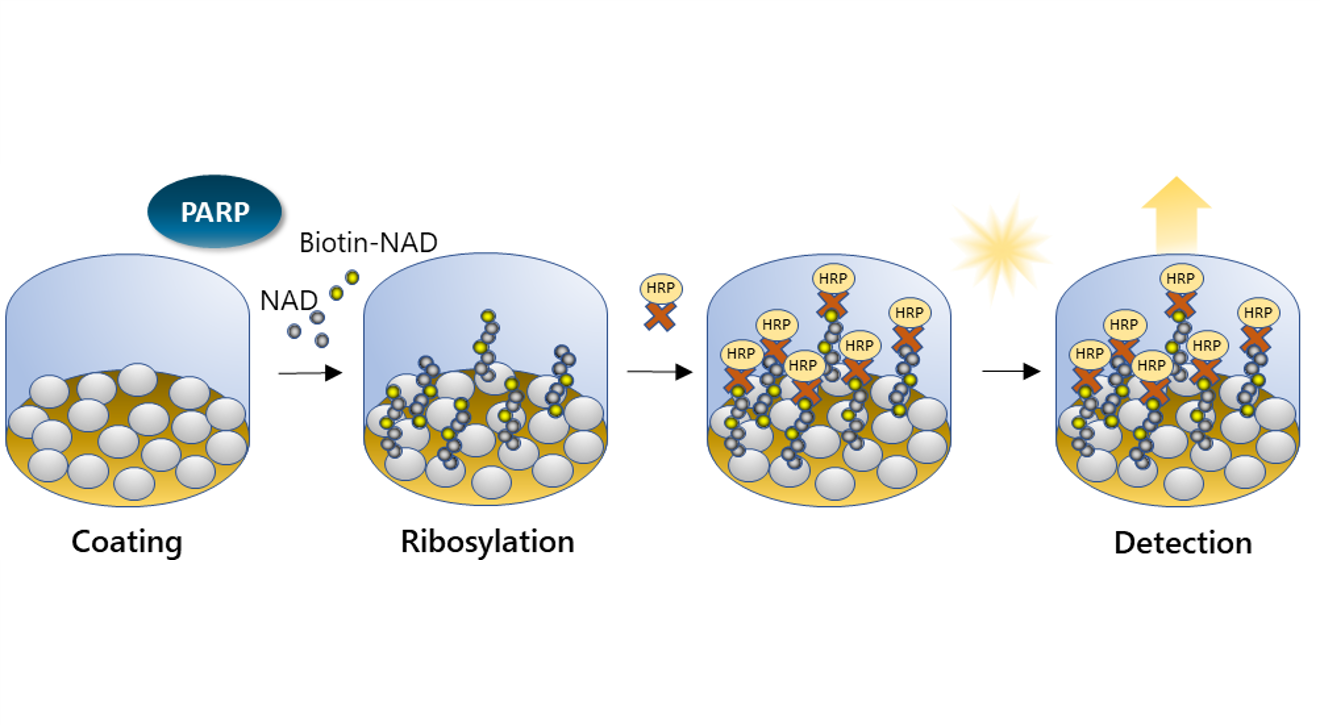
ELISA: Histone proteins are coated on a plate. Biotin-labeled NAD+ is added with the PARP enzyme. PARP-mediated ribosylation activity is detected by adding Streptavidin-HRP (horseradish peroxidase), which binds to the newly formed ribosylation branch, and a chemiluminescent or colorimetric HRP substrate.
Enzymatic Activity by AlphaLISA®
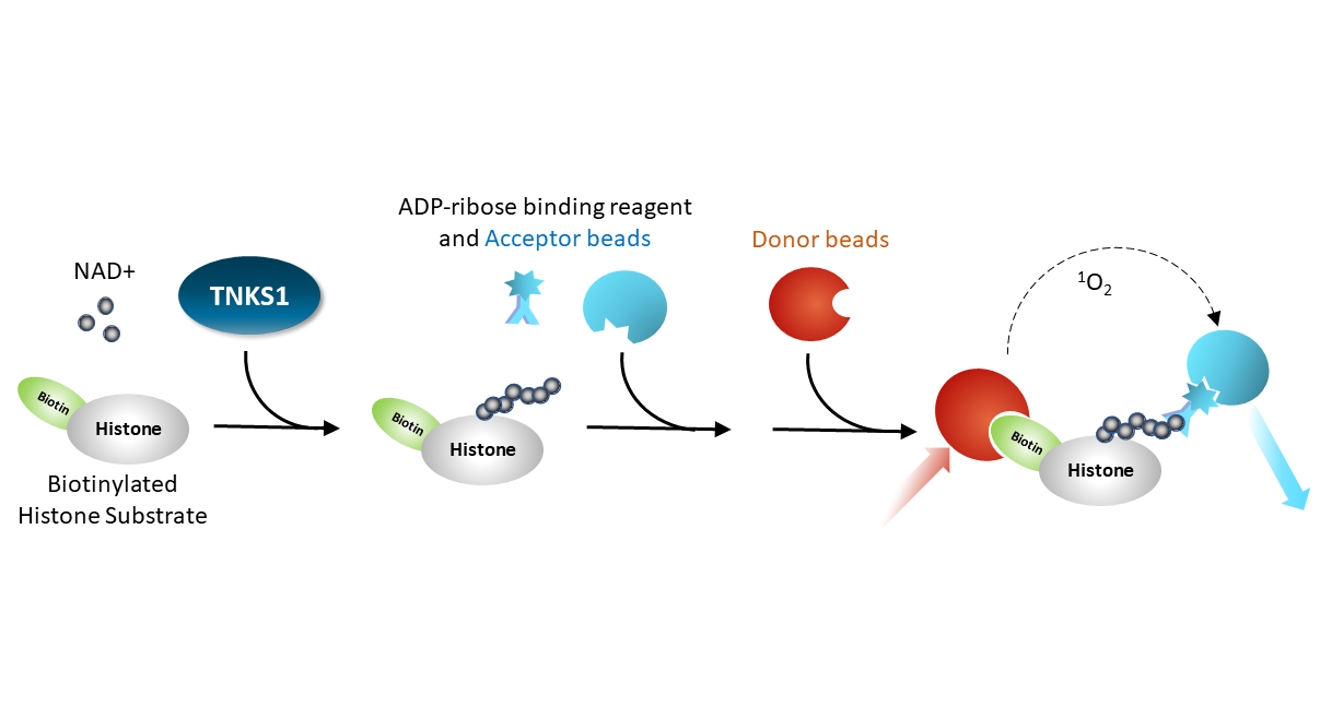
AlphaLISA® Assay: a PARP enzyme is incubated with biotinylated histone substrate and NAD+ for an hour. An ADP-ribose binding reagent is added with an acceptor bead, then a streptavidin-conjugated donor bead is added. Excitation of the donor bead results in excitation of the acceptor bead and light emission. This is a homogeneous assay.
PARP Trapping
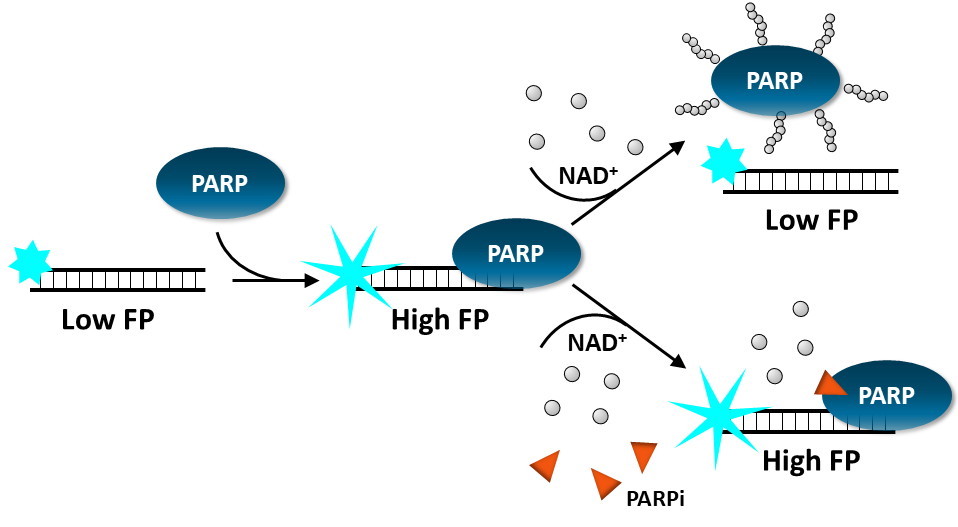
Fluorescence Polarization Assay: The assay uses fluorescent DNA probes that emit differently polarized light depending on PARP binding. Addition of a PARP inhibitor prevents ribosylation and results in the trapping of PARP onto the fluorescent oligonucleotide duplex, which increases the Fluorescence Polarization signal in a dose dependent manner. This is a homogeneous assay.
Molecular Degrader Optimization
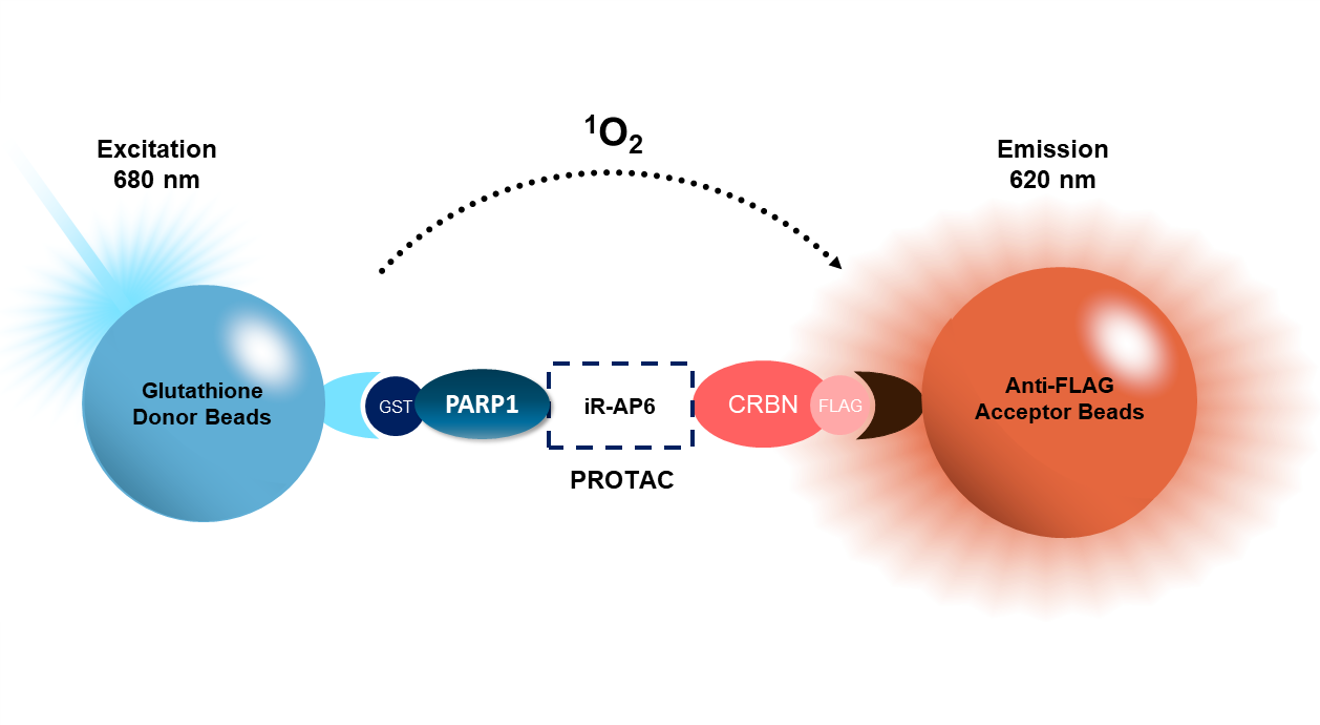
AlphaLISA® Assay: A degrader of interest interacts with both PARP and CRBN, bringing them in proximity. PARP-GST is recognized by the GSH donor bead, while CRBN-FLAG binds to anti-FLAG conjugated to the acceptor bead. Excitation of the donor bead results in excitation of the acceptor bead and light emission. This is a homogeneous assay.

