PARP3 Chemiluminescent Assay Kit
The PARP3 Chemiluminescent Assay Kit is designed to measure the activity of PARP3 (poly-(ADP-ribose) polymerase 3) for screening and profiling applications. The PARP3 assay kit comes in a convenient 96-well or 384-well format, with enough recombinant purified PARP3 enzyme, reaction substrates and PARP assay buffer for 100 or 400 enzyme reactions.
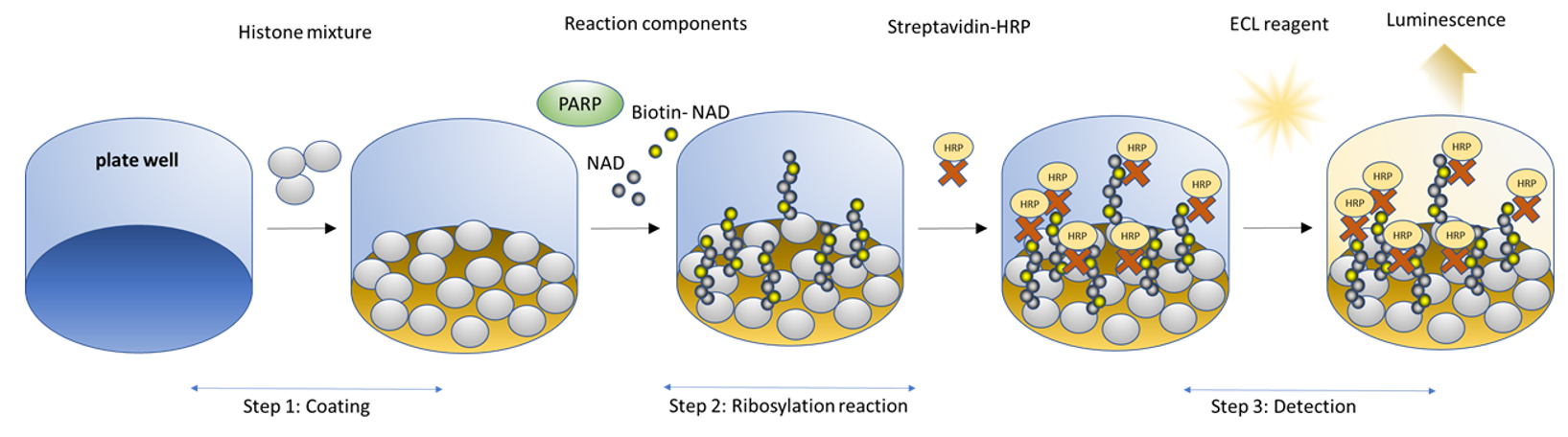
Figure 1. PARP3 Chemiluminescent Assay Kit schematic.
Histone proteins are coated on a 96-well plate. Next, a biotinylated NAD+ mix (termed PARP Substrate Mixture) is incubated with the PARP3 enzyme and an activated DNA template in an optimized assay buffer. The plate is then treated with streptavidin-HRP followed by addition of the ELISA ECL substrate to produce chemiluminescence that can be measured using a chemiluminescence reader. The chemiluminescence signal is proportional to the PARP3 activity.
Need us to run inhibitor screens or profile your compounds against PARP3? Check out our PARP/PARPTrap™ Screening Services or DNA Replication and Repair Screening Services.
This product has been cited 4 times.
- 1x PBS (Phosphate Buffer Saline) Buffer
- PBST Buffer (1x PBS, containing 0.05% Tween-20)
- Luminometer or microplate reader capable of reading chemiluminescence
- Adjustable micropipettor and sterile tips
- Rotating or rocker platform
96 reactions
| Catalog # | Name | Amount | Storage |
| 80503 | PARP3, GST-Tag* | 2 x 1.25 µg | -80°C |
| 52029 | 5x Histone Mixture | 1 ml | -80°C |
| 78366 | PARP Substrate Mixture 1 | 4 x 250 µl | -80°C |
| 80602 | 10x PARP Assay Buffer | 1 ml | -20°C |
| 79743 | Blocking Buffer 3 | 25 ml | +4°C |
| 80605 | Activated DNA | 20 µl | -80°C |
| 0.5 M DTT | 200 µl | -20°C | |
| 80611 | Streptavidin-HRP | 100 µl | +4°C |
| 79670 | ELISA ECL Substrate A (translucent bottle) | 6 ml | Room Temp |
| ELISA ECL Substrate B (brown bottle) | 6 ml | Room Temp | |
| 79837 | 96-well module plate | 1 | Room Temp |
*The concentration of the protein is lot-specific and will be indicated on the tube.
384 reactions
| Catalog # | Name | Amount | Storage |
| 80503 | PARP3, GST-Tag* | 4.5 µg | -80°C |
| 52029 | 5x Histone Mixture | 2 x 1 ml | -80°C |
| 78366 | PARP Substrate Mixture 1 | 8 x 250 µl | -80°C |
| 80602 | 10x PARP Assay Buffer | 2 x 1 ml | -20°C |
| 79743 | Blocking Buffer 3 | 2 x 25 ml | +4°C |
| 80605 | Activated DNA | 2 x 20 µl | -80°C |
| 0.5 M DTT | 2 x 200 µl | -20°C | |
| 80611 | Streptavidin-HRP | 2 x 100 µl | +4°C |
| 79670 | ELISA ECL Substrate A (translucent bottle) | 2 x 6 ml | Room Temp |
| ELISA ECL Substrate B (brown bottle) | 2 x 6 ml | Room Temp | |
| 78188 | 384-well white plate | 1 | Room Temp |
*The concentration of the protein is lot-specific and will be indicated on the tube.
PARP3, also known as poly-(ADP-ribose) polymerase 3 or NAD+ ADP-ribosyltransferase 3, is part of the PARP family. ADP ribosylation, which is the addition of an ADP-ribose to a protein, is a reversible post-translational modification of proteins mostly involved in the DNA Damage Response (DDR) pathway. Mono-ADP-ribosylation (termed MARylation) is the addition of a unit of ADP-ribose. PARP3 is involved in programmed and stress-induced double strand break repair via classical NHEJ (non-homologous end-joining). In the absence of PARP3, DNA repair is less efficient, and cytotoxicity is potentiated when DNA damaging insults occur. It has been linked to cancer, with upregulation of PARP3 occurring in TGFβ (transforming growth factor β)-induced cell EMT (epithelial to mesenchymal transition) in breast cancer. It also promotes cell stemness. The use of inhibitors towards PARP3 can refrain cells from EMT. In addition, the mTORC2 (mTOR complex 2) signaling pathway is impacted, contributing to synthetic lethality in cells deficient in a complementary DNA repair pathway, such as in BRCA1 (breast cancer type 1 susceptibility protein)- triple negative breast cancers (TNBC). PARP3 acts in synergy with PARP1 in the response to ionizing radiation, and the use of inhibitors for both PARP1 and PARP3 also provides an opportunity to reduce the concentration of both and decrease the radiation regimen.
Rodriguez-Vargas J., et al., 2019 Cell Cyle 18(12): 1295-1301